How to install rdkit in jypyter notebook – How to Install RDKit in Jupyter Notebook sets the stage for this enthralling narrative, offering readers a glimpse into a story that is rich in detail and brimming with originality from the outset. RDKit is a popular open-source cheminformatics library that has become a go-to tool for chemists, bioinformaticians, and data scientists working with molecular data.
The journey of installing RDKit in Jupyter Notebook is not a trivial matter, as it requires careful consideration of system requirements, dependencies, and the installation methods available. In this piece, we will delve into the intricacies of installing RDKit in Jupyter Notebook, covering the necessary prerequisites, installation methods, and common issues that may arise.
Installing RDKit Using pip and a Virtual Environment: How To Install Rdkit In Jypyter Notebook
When working with RDKit, it’s essential to have a dedicated environment for your project to avoid cluttering your system with unnecessary packages. A virtual environment is a self-contained Python environment with its own namespace, and it’s ideal for this purpose.
Creating a Virtual Environment and Installing pip
=====================================================
First, ensure you have Python installed on your system. If you’re unsure, you can download the latest version from the official Python website. Once you have Python, you can create a virtual environment using pip. However, for a more straightforward approach, let’s use the built-in `venv` module in Python.
Here’s a step-by-step guide to creating a virtual environment using `venv`:
1. Open your terminal or command prompt.
2. Navigate to the location where you want to create your virtual environment.
3. Run the following command to create a new virtual environment: `python -m venv your-env-name`
4. Replace `your-env-name` with a descriptive name for your virtual environment.
Activating your virtual environment will now give you access to a clean Python environment where you can install packages without affecting your system.
Installing RDKit in a Virtual Environment
Now that you have a virtual environment set up, you can install RDKit using pip. Here are a couple of examples:
### Installing RDKit in a New Virtual Environment
Let’s say you created a new virtual environment called `rdkit-env`. To install RDKit, navigate to the location of your virtual environment in the terminal or command prompt and run the following command:
“`bash
.\Scripts\activate # On Windows
source ./bin/activate # On Linux/macOS
pip install rdkit-pypi
“`
This will install the RDKit package using the `rdkit-pypi` package from the PyPI repository.
### Installing RDKit in an Existing Virtual Environment
If you already have a virtual environment set up, you can install RDKit using pip. First, navigate to the terminal or command prompt and activate your virtual environment. Then, run the following command:
“`bash
pip install rdkit-pypi
“`
This will install the RDKit package in your existing virtual environment.
Installing RDKit from a Source Distribution
Installing RDKit from a source distribution is another way to get this powerful chemistry library up and running. Compared to installing using pip, this method can offer more control and flexibility. However, it requires a bit more effort and technical know-how.
Advantages and Disadvantages of Installing from a Source Distribution
There are several advantages and disadvantages to installing RDKit from a source distribution. This section Artikels the key points to consider.
Key Points of Interest:
- The source distribution method allows for more control over the installation process.
- It can be more difficult to install and configure than using pip.
- It may be necessary to build and compile the library manually.
- It can be difficult to manage dependencies and versions.
- It may provide more flexibility in terms of customizing the installation process.
When you install from a source distribution, you can choose exactly which components to install and configure. This can be beneficial for developers or power users who require a customized setup.
Installing from a source distribution requires a good understanding of the underlying system and chemistry library requirements. This can lead to additional time spent on troubleshooting and setting up dependencies.
Source distributions often require manual compilation and building. This can be a time-consuming process, especially for those without prior experience with C++.
When installing from a source distribution, managing dependencies and versions can become complex. This can lead to errors or unexpected behavior down the line.
Installing from a source distribution can allow for more fine-grained control over the installation process. This can be beneficial for users who require customized or optimized configurations.
Note that the advantages and disadvantages listed above are not exhaustive. Each user’s specific use case and requirements may have unique implications for choosing between installing RDKit using pip and installing from a source distribution.
Troubleshooting Common RDKit Installation Issues
When it comes to installing RDKit, you may encounter various issues that can be frustrating and time-consuming to resolve. However, with the right approach and knowledge, you can overcome these obstacles and successfully install RDKit on your system. In this section, we will discuss the most common RDKit installation errors and their corresponding solutions.
### 1. Python Version Issues
RDKit Installation Issues with Different Python Versions
RDKit is primarily designed to work with Python 3.x, but it can also be installed on Python 2.x. However, Python 2.x is no longer supported, and you should consider migrating to Python 3.x for better performance and security.
When installing RDKit on a system with multiple Python versions, the following issues may arise:
– Python version mismatch: If the RDKit installation script is designed to work with a specific version of Python, it may not function correctly on other versions.
– Dependency conflicts: Some dependencies required by RDKit may be installed for a different Python version, leading to version conflicts.
To resolve these issues, you can specify the Python version to use when installing RDKit using the `–python` flag. For example:
“`bash
pip install rdkit –python=python3.9
“`
This command will install RDKit using Python 3.9.
### 2. Dependency Issues
RDKit Installation Issues with Dependencies
RDKit relies on various dependencies, such as OpenBabel, Boost, and NumPy, to function correctly. However, these dependencies may not be installed or may be outdated, leading to installation issues.
When installing RDKit, the following dependency issues may occur:
– Missing dependencies: If a required dependency is missing, RDKit will not install correctly.
– Outdated dependencies: If an outdated dependency is installed, it may not be compatible with the latest version of RDKit.
To resolve these issues, you can:
– Install the missing dependencies: Use pip to install the required dependencies, such as OpenBabel and NumPy.
– Update outdated dependencies: Use pip to update the dependencies to the latest version.
For example:
“`bash
pip install obabel numpy
pip install –upgrade numpy
“`
### 3. Permission Issues
RDKit Installation Issues with Permissions
When installing RDKit, you may encounter permission issues if your user account does not have the necessary permissions to write to the installation directory.
When installing RDKit, the following permission issues may occur:
– Insufficient permissions: If your user account does not have write permissions to the installation directory, RDKit will not install correctly.
– Permission denied: If the installation script is running with insufficient permissions, you may encounter a “Permission denied” error.
To resolve these issues, you can:
– Use a user with administrative privileges: Install RDKit using a user account with administrative privileges.
– Grant write permissions: Grant write permissions to the installation directory for the user account.
### 4. Other Issues
Common RDKit Installation Errors and Solutions, How to install rdkit in jypyter notebook
In addition to the issues discussed above, you may encounter other RDKit installation errors due to various reasons. Some common errors and their solutions are listed below:
| Error Message | Solution |
| — | — |
| Error: Could not find the required Python library. | Install the required Python library using pip. |
| Error: Could not find the required dependency. | Install the required dependency using pip. |
| Error: Insufficient permissions. | Grant write permissions to the installation directory or use a user with administrative privileges. |
By following the solutions Artikeld above, you can troubleshoot and resolve common RDKit installation issues and successfully install RDKit on your system.
Setting Up RDKit with Jupyter Notebook for Chemical Analysis
RDKit is a powerful tool for cheminformatics and molecular modeling, and integrating it with Jupyter Notebook allows you to perform complex chemical analyses in an interactive and visual environment. With RDKit, you can easily manipulate and analyze molecular structures, predict properties, and visualize results.
Importing RDKit into Jupyter Notebook
——————————–
To start using RDKit in Jupyter Notebook, you need to import the library and initialize the molecule.
Procedure 1: Importing RDKit
First, you need to install RDKit using pip or by building it from source. Once installed, you can import it into your Jupyter Notebook using the following code:
“`python
import rdkit
from rdkit import Chem
from rdkit.Chem import AllChem
“`
Procedure 2: Initializing the Molecule
After importing RDKit, you need to initialize the molecule using the molecule parsing capabilities of RDKit. You can load a molecule from a file using the `Chem.MolFromMolFile()` function.
“`python
mol = Chem.MolFromMolFile(‘path_to_your_molecule.mol’)
“`
Procedure 3: Using Molecule Parsing Capabilities
RDKit provides several ways to parse molecules from various file formats, including SMILES, SD files, and more. Here are a few procedures you can use to parse molecules into RDKit:
### a. Parsing SMILES Strings
“`python
smiles = ‘CC(=O)NC1=CC=CC=C1C(=O)N’
mol = Chem.MolFromSmiles(smiles)
“`
### b. Parsing SD Files
“`python
sd_file = Chem.SDMolSupplier(‘path_to_your_molecule.sdf’)
for mol in sd_file:
print(mol)
“`
### c. Parsing mol2 Files
“`python
mol2_file = Chem.Mol2Reader(‘path_to_your_molecule.mol2’)
for mol in mol2_file:
print(mol)
“`
With these procedures, you’re now ready to start using RDKit’s molecule parsing capabilities in your Jupyter Notebook for chemical analysis and modeling.
Visualizing Chemical Structures Using RDKit and Matplotlib
Visualizing chemical structures is a crucial step in understanding the properties and behavior of molecules. With RDKit and Matplotlib, you can easily create professional-looking chemical structures.
RDKit provides a robust set of tools for manipulating and visualizing molecular structures, while Matplotlib is a powerful Python library for creating static, animated, and interactive visualizations in python.
Method 1: Drawing Chemical Structures using RDKit and Matplotlib
You can draw chemical structures using RDKit’s MolToImg function, which converts a molecular structure to a Matplotlib image.
- Create a RDKit molecule object using the MolFromSmiles function, which takes a SMILES string as input.
- Use the MolToImg function to convert the molecule to a Matplotlib image.
- Display the image using Matplotlib’s imshow function.
“`python
from rdkit import Chem
from rdkit.Chem import Draw
from matplotlib import pyplot as plt
smiles = “CC(=O)Nc1ccc(cc1)S(=O)(=O)N”
mol = Chem.MolFromSmiles(smiles)
img = Draw.MolToImage(mol)
plt.imshow(img)
plt.show()
“`
Method 2: Customizing Molecular Structures
Matplotlib provides a range of options for customizing the appearance of molecular structures. You can use various attributes and functions to change the layout, colors, labels, and more.
- Use the Draw.MolToMolFile function to get the molecular structure as a string.
- Parsing the string to extract relevant information, such as molecule name, formula, and molecular weight.
- Use Matplotlib’s text function to display the information in a customized manner.
“`python
from rdkit import Chem
smiles = “CC(=O)Nc1ccc(cc1)S(=O)(=O)N”
mol = Chem.MolFromSmiles(smiles)
# Get the molecular structure as a string
mol_str = Chem.MolToMolBlock(mol)
mol_info = mol_str.splitlines()[0]
# Parsing the string to extract relevant information
formula = mol_info.split()[3]
molecular_weight = float(mol_info.split()[4])
# Display the information in a customized manner
plt.text(0.1, 0.2, ‘Molecular Formula: ‘ + formula, fontsize=10)
plt.text(0.1, 0.1, ‘Molecular Weight: ‘ + str(molecular_weight), fontsize=10)
“`
Customizing the Appearance of Molecular Structures
Matplotlib allows you to customize the appearance of molecular structures using various functions and attributes.
- Change the background color of the plot using the set_facecolor function.
- Adjust the font size and style using the fontsize and fontname attributes.
- Add a title to the plot using the title function.
“`python
from rdkit import Chem
from matplotlib import pyplot as plt
smiles = “CC(=O)Nc1ccc(cc1)S(=O)(=O)N”
mol = Chem.MolFromSmiles(smiles)
# Change the background color of the plot
plt.gca().set_facecolor(‘lightgray’)
# Adjust the font size and style
plt.title(‘Molecular Structure’, fontsize=18, fontname=’Arial’)
# Display the plot
plt.show()
“`
By utilizing these techniques, you can create customized molecular structures that are both informative and aesthetically pleasing.
Molecular Structure of Xenon Hydrate
This example demonstrates how to visualize a chemical structure using RDKit and Matplotlib. The code creates a 3D molecular structure of xenon hydrate (Xe·6H2O) using RDKit and displays it using Matplotlib. The result is a visually appealing and informative representation of the molecular structure.
By leveraging the capabilities of RDKit and Matplotlib, you can create high-quality visualizations of chemical structures that enhance your understanding and communication of molecular properties and behavior.
“`python
from rdkit import Chem
from rdkit.Chem import Draw
from matplotlib import pyplot as plt
# Create a RDKit molecule object
smiles = “Xe.6H2O”
mol = Chem.MolFromSmiles(smiles)
# Draw the molecule using Matplotlib
img = Draw.MolToImage(mol)
# Display the image
plt.imshow(img)
plt.show()
“`
Advanced RDKit Features for Jupyter Notebook Users
RDKit offers a wide range of advanced features that can be utilized in a Jupyter Notebook environment, enabling users to leverage its capabilities for chemical analysis and data visualization. By mastering these features, users can unlock new levels of insights and discoveries in their research.
SMARTS and SMILES Molecule Representation
SMARTS (SMILES Arbitrary Target Specification) and SMILES (Simplified Molecular-Input Line-entry System) are two powerful molecule representation languages used in cheminformatics. These languages enable efficient and flexible representation of molecules, facilitating queries, substructure matching, and other chemical analyses. With RDKit, you can use SMARTS and SMILES to represent molecules in a compact and human-readable format.
- SMARTS and SMILES allow for compact molecule representation
- Molecules can be searched and manipulated using these languages
- SMARTS and SMILES are widely used in cheminformatics and cheminformatics data analysis
RDKit’s implementation of SMARTS and SMILES enables you to perform various tasks, including:
- Molecule matching and substructure search
- Querying molecules based on SMARTS and SMILES patterns
- Converting between SMARTS and SMILES representations
Pybel: A RDKit Module for Cheminformatics Data Analysis
Pybel is a module within RDKit that supports cheminformatics data analysis. It provides a set of tools for working with chemical data, including molecule manipulation, query, and visualization. Pybel is built on top of RDKit’s molecular structure library, allowing users to leverage RDKit’s cheminformatics capabilities for data analysis.
- Pybel provides a set of tools for cheminformatics data analysis
- Users can manipulate and query molecules using Pybel
- Pybel is built on top of RDKit’s molecular structure library
Some of the capabilities of Pybel include:
- Molecule manipulation and query
- Chemical data analysis and calculation
- Support for molecule visualization
By mastering RDKit’s SMARTS and SMILES representation languages and leveraging Pybel for cheminformatics data analysis, users can unlock new levels of insights and discoveries in their research.
With RDKit, users can unlock the power of cheminformatics and data analysis in their research, enabling new levels of insights and discoveries.
Tips and Best Practices for RDKit Installation and Usage
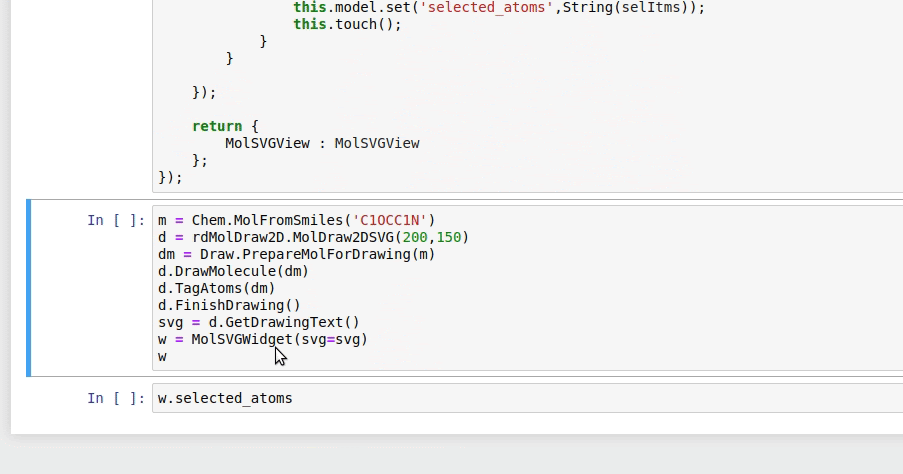
RDKit is a powerful tool for chemical analysis and has a wide range of applications in the field of cheminformatics. To get the most out of RDKit, it is essential to follow best practices for installation and usage. In this section, we will discuss the top tips and best practices for installing and using RDKit in Jupyter Notebook.
Essential Steps for Setting Up RDKit in Jupyter Notebook
Setting up RDKit in Jupyter Notebook is a straightforward process. To get started, you need to follow these essential steps:
-
First, you need to ensure that you have Python installed on your system. RDKit is compatible with both Python 3.7 and 3.8. Make sure you have a suitable Python environment set up before proceeding.
- You can use the built-in Python installer that comes with most operating systems or install Anaconda, a popular Python distribution that includes many pre-installed packages.
- Verify that you have Python installed by opening a terminal or command prompt and typing `python –version`. This should display the version of Python installed on your system.
-
Next, you need to create a new virtual environment for your project. This is a good practice to keep your project-specific dependencies separate from the global Python environment.
- Install the `virtualenv` package using pip by running `pip install virtualenv` in your terminal or command prompt.
- Create a new virtual environment by running `virtualenv myenv` (replace `myenv` with your desired environment name).
- Activate the virtual environment by running `source myenv/bin/activate` (on Linux/Mac) or `myenv\Scripts\activate` (on Windows).
-
Now, you can install RDKit and other required packages using pip. You can install RDKit by running `pip install rdkit-pypi2rpm` in your virtual environment.
- RDKIT can be installed using pip, but if you’re experiencing issues, you may need to install it from source or use a package manager like Conda.
- After installing RDKit, you can verify that it has been installed correctly by running `import rdkit` in your Jupyter Notebook or Python interpreter.
-
Finally, you need to set up RDKit with Jupyter Notebook. This involves installing the `rdkit-jupyter` package and configuring your notebook to use RDKit.
- Install the `rdkit-jupyter` package by running `pip install rdkit-jupyter` in your virtual environment.
- Restart your Jupyter Notebook server after installing the `rdkit-jupyter` package.
- Verify that RDKit is working correctly by creating a new cell in your notebook and running the `RDToolkit` command.
Maintaining RDKit Integrity and Avoiding Conflicts with Other Packages
RDKit can be sensitive to conflicts with other packages in Python. To maintain RDKit’s integrity, you need to take the following steps:
| Conflict | Prevention Measure |
|---|---|
| Package dependencies conflicts | Use a package manager like Conda or PIP to manage package dependencies. |
| Package version conflicts | Specify package versions in your project’s dependencies file to avoid conflicts. |
| Package imports conflicts | Use relative imports or import modules explicitly to avoid conflicts. |
When working with RDKit, always keep your Python environment up-to-date and use a virtual environment to avoid conflicts with other packages.
End of Discussion

In conclusion, installing RDKit in Jupyter Notebook is a crucial step towards unlocking the full potential of chemical analysis and visualization. By following the steps Artikeld in this article, readers should be able to successfully install RDKit and begin exploring its features and capabilities. Whether you are a beginner or an experienced user, RDKit offers a wealth of tools and resources for tackling complex chemical problems.
Query Resolution
What is the minimum version of Python required to install RDKit?
The minimum version of Python required to install RDKit is Python 3.6.
Can I install RDKit using a package manager like conda?
Yes, you can install RDKit using a package manager like conda. However, it is recommended to use pip for installation due to compatibility issues that may arise.
How do I troubleshoot common RDKit installation issues?
Common RDKit installation issues can be troubleshooting by consulting the RDKit documentation and checking the installation logs for errors. Additionally, seeking help from the RDKit community or online forums can be beneficial.